HydraLA-Net: Multi-Lesion Segmentation for Diabetic Retinopathy Detection
Introduction
Diabetic retinopathy (DR) is one of the leading causes of preventable blindness worldwide, affecting millions of people with diabetes. Early detection and treatment can prevent up to 95% of vision loss cases, but manual screening is time-consuming and requires expert ophthalmologists.
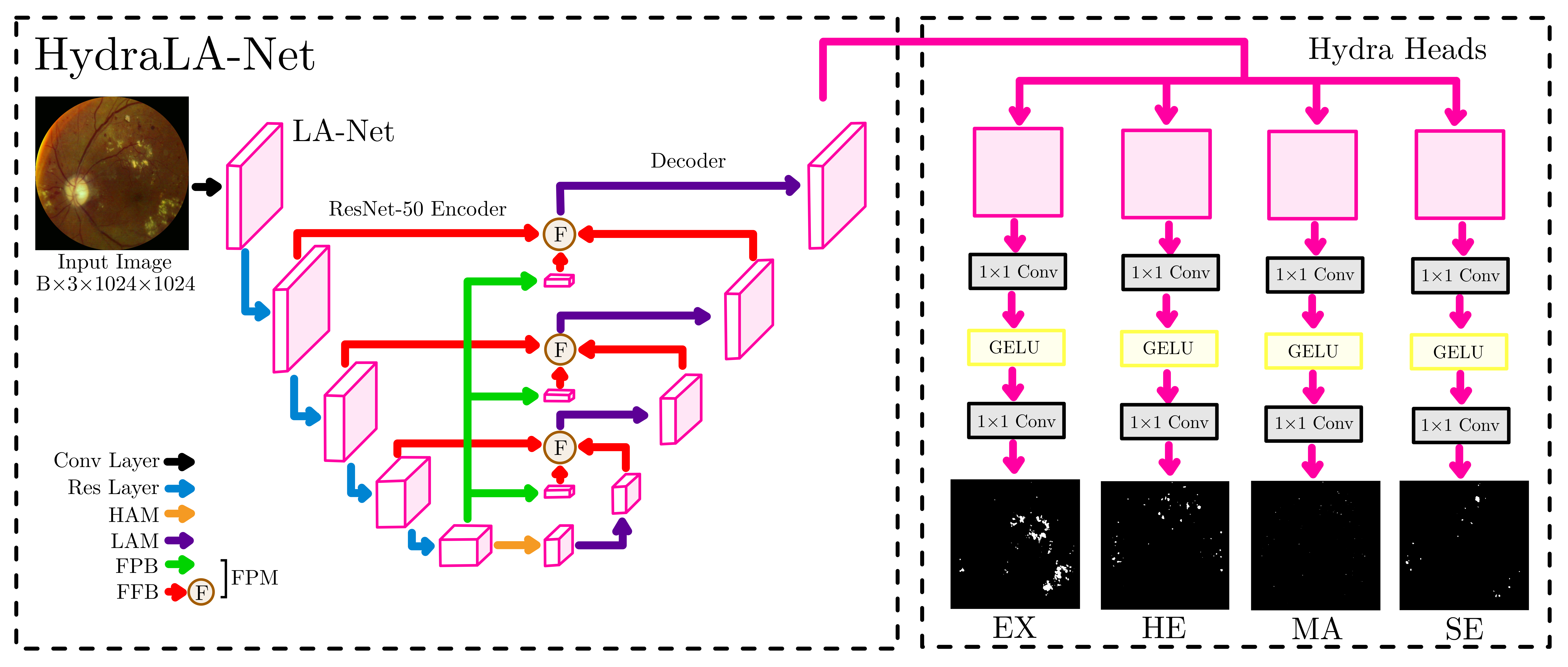
In this project, I developed HydraLA-Net, a deep learning system that automatically segments four types of retinal lesions in fundus images: Hard Exudates (EX), Hemorrhages (HE), Microaneurysms (MA), and Soft Exudates (SE). The system is built on the LA-Net (Lesion-Aware Network) architecture with several novel enhancements, particularly a "Hydra" segmentation head that improves per-class performance.
The Challenge: Tiny Lesions in Low-Contrast Images
Microaneurysms are particularly challenging to detect - they're often just a few pixels wide and appear as subtle red dots on the retina. The dataset suffers from severe class imbalance: microaneurysms comprise only a tiny fraction of pixels compared to the background.
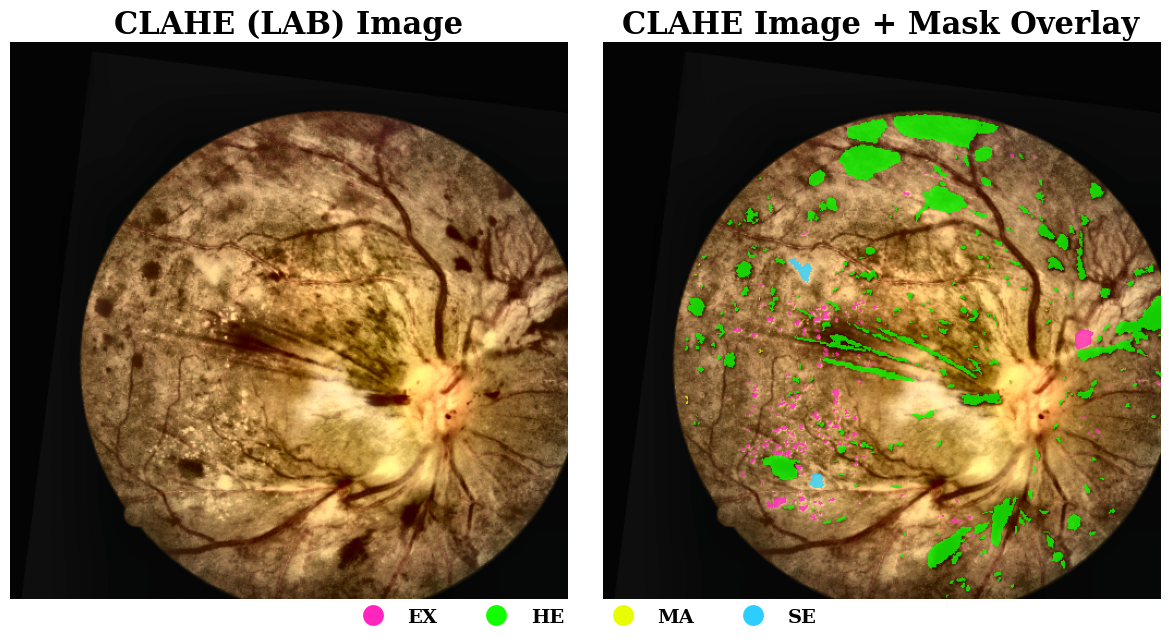
Traditional approaches struggle with this imbalance, often missing small lesions entirely or generating too many false positives.
Part 1: Data Preprocessing - Why CLAHE and Stratified Splitting Matter
Dataset Aggregation
I combined three publicly available datasets to improve model generalization:
- IDRiD (81 images): High-quality Indian dataset with binary masks
- DDR (757 images): Large-scale dataset with .tif format masks
- TJDR (561 images): Color-coded annotations requiring parsing
Total: 1,399 images split into:
- Training: 827 samples (59%)
- Validation: 207 samples (15%)
- Test: 365 samples (26%)
Why Stratified Splitting?
Here's where the data pipeline gets interesting. Instead of random splitting, I used stratified splitting to ensure balanced class representation across splits.
Random splitting would randomly assign images to train/val/test sets, which could result in:
- Some classes being over-represented in training but under-represented in validation
- Rare lesions (like microaneurysms) potentially missing from certain splits
- Inconsistent difficulty across splits
Stratified splitting ensures each split maintains the same class distribution as the original dataset. This is crucial for medical imaging where certain lesion types are already rare.
Here's a snippet from my data pipeline:
from sklearn.model_selection import train_test_split
# First split: separate out test set (26%)
train_val_df, test_df = train_test_split(
combined_df,
test_size=0.26,
random_state=42,
stratify=combined_df['dataset'] # Stratify by dataset source
)
# Second split: separate train and validation (59% / 15% of total)
train_df, val_df = train_test_split(
train_val_df,
test_size=0.2, # 20% of 74% = 15% of total
random_state=42,
stratify=train_val_df['dataset']
)This ensures each dataset (IDRiD, DDR, TJDR) is proportionally represented in all splits, preventing the model from overfitting to characteristics of a single data source.
CLAHE: Enhancing Contrast for Better Lesion Visibility
Fundus images often have low contrast, making tiny lesions difficult to detect. I implemented CLAHE (Contrast Limited Adaptive Histogram Equalization) to enhance local contrast without over-amplifying noise.
Why CLAHE instead of standard histogram equalization?
Standard histogram equalization operates globally on the entire image, which can:
- Over-amplify noise in already bright/dark regions
- Create unnatural-looking artifacts
- Reduce diagnostic quality
CLAHE operates on small tiles (8×8 pixels in my case) and limits contrast enhancement via a clip limit (1.5), preventing noise amplification while still revealing subtle lesions.
I tested three CLAHE modes:
1. LAB Mode: Apply CLAHE to the L (lightness) channel in LAB color space 2. Green Channel Mode (selected for final model): Apply CLAHE only to the green channel 3. CASP Mode: Channel-Aware Selective Preprocessing with different processing per channel
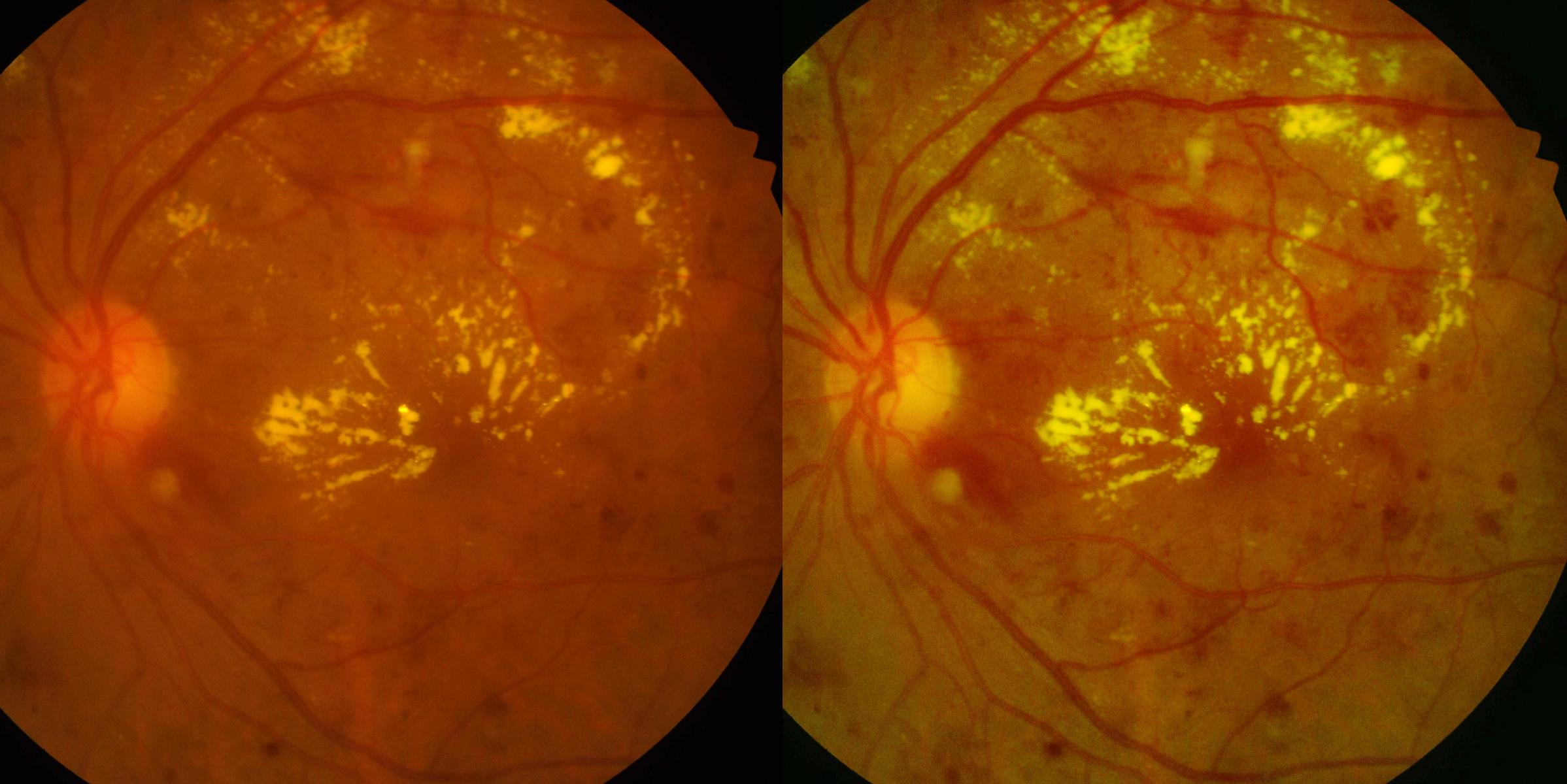
Why did I choose Green Channel CLAHE?
The green channel provides the best contrast for blood vessels and microaneurysms in fundus photography - it's standard practice in retinal imaging. Red channels are often overexposed, and blue channels have poor SNR.
Here's the implementation from dataset_definition.py:
def apply_clahe(
image_rgb, clip_limit = 2.5, tile_grid_size = (8, 8), mode = "casp"
):
clahe = cv2.createCLAHE(
clipLimit = clip_limit,
tileGridSize = tile_grid_size
)
if mode == "lab":
lab = cv2.cvtColor(image_rgb, cv2.COLOR_RGB2LAB)
l, a, b = cv2.split(lab)
l = clahe.apply(l)
lab = cv2.merge((l, a, b))
return cv2.cvtColor(lab, cv2.COLOR_LAB2RGB)
elif mode == "green":
out = image_rgb.copy()
out[:, :, 1] = clahe.apply(out[:, :, 1])
return outPart 2: Model Architecture - Building HydraLA-Net
Base Architecture: LA-Net (Lesion-Aware Network)
I started with LA-Net, which uses:
- ResNet-50 backbone (pretrained on ImageNet) for feature extraction
- Feature Pyramid Blocks (FPB) to create multi-scale representations
- Lesion-Aware Modules (LAM) with asymmetric convolutions to capture elongated lesion shapes
- Feature Fusion Blocks (FFB) to combine encoder and decoder features
Innovation: The Hydra Segmentation Head
The key innovation in HydraLA-Net is the Hydra head - a multi-branch segmentation head where each lesion type gets its own dedicated branch.
Why separate branches?
Traditional multi-class segmentation uses a single convolutional layer to produce all class predictions simultaneously. This creates competition between classes and can hurt performance when classes have vastly different characteristics (e.g., large hemorrhages vs. tiny microaneurysms).
The Hydra head has 4 independent branches (hydralanet.py:14-33):
class HydraSegHead(nn.Module):
def __init__(self, in_ch = 256, hidden_ratio = 0.25):
super().__init__()
hidden = max(1, int(in_ch * hidden_ratio))
self.heads = nn.ModuleDict({
name: nn.Sequential(
nn.Conv2d(in_ch, hidden, kernel_size = 1, bias = True),
nn.GELU(),
nn.Conv2d(hidden, 1, kernel_size = 1, bias = True),
)
for name in ["EX", "HE", "MA", "SE"]
})
for head in self.heads.values():
nn.init.constant_(head[-1].bias, -2.0)
def forward(self, x):
logits = [head(x) for head in self.heads.values()]
return torch.cat(logits, dim = 1)Key design choices:
- Hidden ratio of 0.25: Reduces parameters (256→64→1) while maintaining expressiveness
- GELU activation: Smoother gradients than ReLU, better for small lesions
- Bias initialization to -2.0: With sigmoid activation, this starts predictions near 0.12, reducing false positives early in training
Each branch learns specialized features for its lesion type, avoiding the compromises of a shared prediction layer.
Part 3: Handling Class Imbalance with Dual Loss
The biggest challenge in this project was severe class imbalance. Microaneurysms might occupy less than 0.01% of image pixels, while the background dominates.
![]()
I implemented a Dual Loss combining Focal Tversky Loss and Binary Cross-Entropy:
Why Focal Tversky Loss?
Tversky Index is a generalization of Dice/F1 that allows asymmetric weighting of false positives and false negatives:
Tversky = TP / (TP + α×FP + β×FN)- α (FP penalty): How much we penalize false positives
- β (FN penalty): How much we penalize false negatives
For medical imaging, false negatives (missed lesions) are more dangerous than false positives, so β > α.
Here's the actual implementation from loss_functions.py:16-92:
class FocalTverskyLoss(nn.Module):
"""
Multi-Label Focal Tversky Loss for Semantic Segmentation
alpha: controls penalty on false positives
beta: controls penalty on false negatives
gamma: focal parameter
smooth: used to prevent division by 0
class_weights: explicit scalar reweighting per class
"""
def __init__(
self, alpha = 0.3, beta = 0.7, gamma = 1.3, smooth = 1e-6, class_weights = None,
):
super().__init__()
self.register_buffer("alpha", torch.as_tensor(alpha, dtype = torch.float32))
self.register_buffer("beta", torch.as_tensor(beta, dtype = torch.float32))
self.register_buffer("gamma", torch.as_tensor(gamma, dtype = torch.float32))
if class_weights is None:
class_weights = torch.ones(4)
self.register_buffer(
"class_weights",
torch.as_tensor(class_weights, dtype = torch.float32)
)
self.register_buffer(
"smooth",
torch.tensor(smooth, dtype = torch.float32)
)
def forward(self, probs, targets):
# ... safety checks omitted for brevity ...
# Flatten spatial dims
probs_flat = probs.view(B, C, -1)
targets_flat = targets.view(B, C, -1)
# TP / FP / FN
TP = (probs_flat * targets_flat).sum(dim = 2)
FP = (probs_flat * (1 - targets_flat)).sum(dim = 2)
FN = ((1 - probs_flat) * targets_flat).sum(dim = 2)
# Tversky + focal term
tversky = (TP + self.smooth) / (
TP + alpha * FP + beta * FN + self.smooth
)
loss_per_class = (1 - tversky) ** gamma
weighted_loss = (loss_per_class.mean(dim = 0) * self.class_weights).mean()
return weighted_lossNote: For microaneurysms (MA), I used β=0.8, α=0.2 - heavily penalizing missed microaneurysms while being more tolerant of false positives.
The Dual Loss: Best of Both Worlds
I combined Focal Tversky with Binary Cross-Entropy (loss_functions.py:154-200):
class DualLoss(nn.Module):
"""
Combined Loss Function (Mixed Focal Tversky and BCE)
Final Loss: L = w_ft * FocalTverskyLoss + w_bce * BCELossMultiLabel
"""
def __init__(
self, class_weights = None, w_ft = 0.5, w_bce = 0.5, alpha = 0.3, beta = 0.7,
gamma = 1.3, smooth = 1e-6, pos_weight = None
):
super().__init__()
self.w_ft = w_ft
self.w_bce = w_bce
self.ft = FocalTverskyLoss(
alpha = alpha,
beta = beta,
gamma = gamma,
smooth = smooth,
class_weights = class_weights,
)
self.bce = BCELossMultiLabel(
class_weights = class_weights,
pos_weight = pos_weight,
)
def forward(self, probs, targets):
loss_ft = self.ft(probs, targets)
loss_bce = self.bce(probs, targets)
return self.w_ft * loss_ft + self.w_bce * loss_bceWhy combine them?
- Focal Tversky: Handles class imbalance and focuses on hard examples
- BCE: Provides stable pixel-wise gradients and prevents collapse
With weights 0.8 and 0.2 respectively, plus class weights [1.0, 1.0, 5.0, 1.0], giving 5× importance to microaneurysm predictions.
Part 4: Training Strategy and Results
Training Configuration
From train.py:36-104:
# Global Constants
IMG_SIZE = 1024
DEFAULT_EPOCHS = 125
LEARNING_RATE = 1e-5
BATCH_SIZE = 2
NUM_WORKERS = 8
W_FTL = 0.8
W_BCE = 0.2
TVERSKY_ALPHA = [0.4, 0.4, 0.2, 0.4]
TVERSKY_BETA = [0.6, 0.6, 0.8, 0.6]
TVERSKY_GAMMA = 2
CLAHE_CLIP = 1.5
CLAHE_MODE = 'green'
CLASS_WEIGHTS = [1.0, 1.0, 5.0, 1.0]
THRESHOLDS = [0.35, 0.35, 0.35, 0.35]
# Model initialization
model = HydraLANet()
model = model.to(device)
# Optimizer
optimizer = torch.optim.AdamW(
params = model.parameters(),
lr = LEARNING_RATE,
weight_decay = 1e-4
)
# Loss Function
loss_function = DualLoss(
class_weights = CLASS_WEIGHTS,
w_ft = W_FTL,
w_bce = W_BCE,
alpha = TVERSKY_ALPHA,
beta = TVERSKY_BETA,
gamma = TVERSKY_GAMMA,
smooth = 1e-6
).to(device)Key training decisions:
-
Small batch size (2): Required due to GPU memory with 1024×1024 images. I used frozen BatchNorm layers to avoid instability from batch statistics.
-
AdamW with low learning rate (1e-5): Fine-tuning pretrained ResNet-50 requires careful learning rates to avoid catastrophic forgetting.
-
High resolution (1024×1024): Microaneurysms can be just 3-5 pixels wide. Lower resolutions (512×512) caused significant performance drops.
Data Augmentation
From dataset_definition.py:131-144:
if self.transform_type == "train":
transforms.extend([
A.HorizontalFlip(p = 0.5),
A.VerticalFlip(p = 0.5),
A.Affine(
translate_percent = 0.1,
scale = (0.88, 1.25),
rotate = (-15, 15),
border_mode = cv2.BORDER_CONSTANT,
fill = 0,
fill_mask = 0,
p = 0.75,
),
])Medical images can have any orientation, so rotations and flips are valid augmentations. I avoided color augmentations since CLAHE preprocessing already handles contrast.
Evaluation Metrics
I tracked three metrics per class:
- IoU (Intersection over Union): Segmentation accuracy
- F1 Score: Harmonic mean of precision and recall (primary metric)
- Recall: Critical for medical applications - we don't want to miss lesions
Results are calculated per class and then averaged, giving equal importance to each lesion type despite class imbalance.
Part 5: Deployment with Streamlit
I deployed the model as an interactive web app using Streamlit on Hugging Face Spaces.
Key preprocessing function from app.py:15-38:
def apply_green_clahe(image_rgb, clip_limit=1.5, tile_grid_size=(8, 8)):
"""Apply CLAHE to green channel only for contrast enhancement"""
clahe = cv2.createCLAHE(
clipLimit = clip_limit,
tileGridSize = tile_grid_size
)
out = image_rgb.copy()
out[:, :, 1] = clahe.apply(image_rgb[:, :, 1])
return out
def preprocess_image(image_rgb):
"""Preprocess image with CLAHE and ImageNet normalization"""
image_rgb = apply_green_clahe(image_rgb)
image = image_rgb.astype(np.float32) / 255.0
mean = np.array([0.485, 0.456, 0.406], dtype=np.float32)
std = np.array([0.229, 0.224, 0.225], dtype=np.float32)
image = (image - mean) / std
image_tensor = torch.from_numpy(image).permute(2, 0, 1).unsqueeze(0)
return image_tensor, image_rgbThe app allows users to:
- Upload their own fundus images
- Adjust detection threshold for sensitivity vs. specificity trade-offs
- Toggle CLAHE preprocessing
- View individual masks and coverage statistics
Conclusions and Key Takeaways
This project demonstrates several important principles for medical image segmentation:
1. Preprocessing Matters
CLAHE preprocessing on the green channel significantly improved lesion visibility without introducing artifacts. Domain-specific preprocessing (using medical imaging knowledge) outperformed generic approaches.
2. Architecture Innovations Can Be Simple
The Hydra head required minimal code changes but provided measurable improvements by giving each class its own decision pathway.
3. Loss Functions Are Critical for Imbalance
The combination of Focal Tversky Loss (with class-specific α/β parameters) and class weighting was essential for learning to detect rare microaneurysms.
4. Multi-Dataset Training Improves Generalization
Combining three datasets with different annotation formats and imaging protocols forced the model to learn robust features rather than dataset-specific artifacts.
5. Resolution Cannot Be Compromised
For tiny lesions like microaneurysms, high-resolution training (1024×1024) was non-negotiable. The performance drop at 512×512 was significant.
Impact
Automated diabetic retinopathy screening can help address the shortage of ophthalmologists in underserved areas, enabling earlier detection and treatment. By detecting multiple lesion types simultaneously, systems like HydraLA-Net can provide more comprehensive assessments than single-lesion detectors.
Try It Yourself
- Live Demo: Hugging Face Spaces
- Paper: Progressive Optimization of HydraLA-Net for Microaneurysm Segmentation
- Code: GitHub Repository
Built with PyTorch, Streamlit, and a lot of coffee
Tags: #DeepLearning #MedicalAI #ComputerVision #DiabeticRetinopathy #SemanticSegmentation #PyTorch